Structural Basis of Protein Signaling in Phototransduction Pathways
I. Interactions between proteins of the visual signaling cascade
The process of phototransduction involves absorption of light stimulus by sensor proteins that trigger a number of signaling pathways in nearly all organisms. In vertebrates, the first step of conversion of a photon into an electrical signal takes place in the retina in the eye via a G-protein coupled receptor rhodopsin. Absorption of photon leads to the isomerization of 11-cis to all-trans retinal followed by conformational changes of the protein, which then binds the GTP-binding protein transducin. The subsequent amplification cascades lead to the closure of cyclic GMP-gated cation channel and hyperpolarization of the photoreceptor cell, resulting in a neural impulse.
An essential step in visual signaling is the efficient termination and rapid recovery of the photo-receptor. At molecular level, this begins by inactivation of rhodopsin that involves a two-step mechanism – a rapid phosphorylation, followed by the binding of arrestin (figure). The latter step sterically blocks further interaction of G-protein with the receptor, terminating the primary signaling event.

Our Team focuses on studying structural and biochemical properties of arrestin family members, which play a central role in the termination of G-protein coupled receptor (GPCR) mediated signal transduction pathways. Recently we determined crystal structures of arrestin and its constitutively active splice variant p44. The structural details in combination with the biochemical properties provide a better understanding of the nature of interaction between arrestin and the GPCR. The results are significant as GPCRs are the largest family of signaling proteins in mammalian genomes and are targets for majority of clinically used drugs.
II. Structural studies of blue-light photoreceptors from the class of LOV proteins
In a second project, our team is investigating the structures of blue-light photoreceptors containing light-oxygen-voltage (LOV) domains, which regulate a myriad of different physiological responses in both eukaryotes and prokaryotes. Due to their intrinsic photochemical /photophysical properties, LOV proteins have proven as novel fluorescent reporters and blue-light sensitive photo-switches. Biotechnological applications include their use as real-time oxygen independent flavin-based fluorescent reporter proteins (FbFPs), as biological trap to produce flavin mononucleo-tide (FMN), as well as in development of LOV-based optogenetic tools.
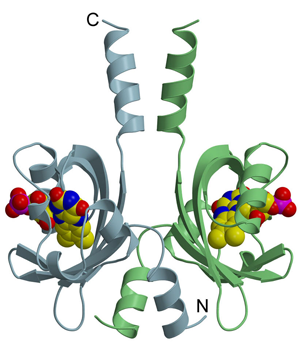
All LOV domains undergo the same primary photochemistry leading to a covalent adduct formation between a conserved cysteine residue and the chromophore FMN; however, considerable variation is found in the lifetime of the adduct state that varies from seconds to several hours. The molecular mechanism underlying this variation among the structurally conserved LOV protein family is not well understood. Based on the crystal structures of LOV proteins studied in the Team, we could explain the very slow dark recovery of a full-length LOV protein from Pseudomonas putida. Moreover, a complementary phylogenetic analysis highlights a common ancestry of bacterial LOV photoreceptors and Per-ARNT-Sim chemosensors.

